L-Lysine (PAMDB000075)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000075 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
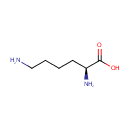
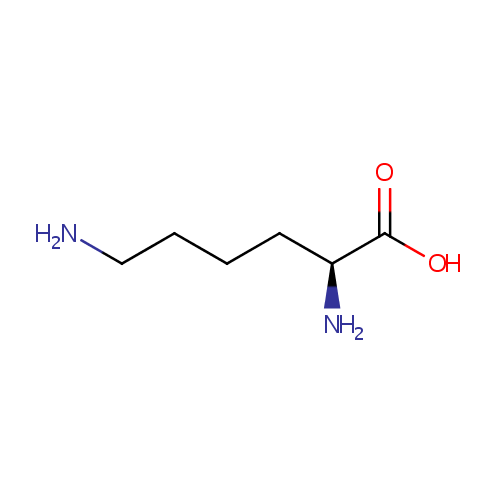
| Name: | L-Lysine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | L-lysine is an alpha-amino acid with the chemical formula HO2CCH(NH2)(CH2)4NH2. Lysine is a basic amino acid as are arginine and histidine. Lysine is a proteogenic amino acid, meaning that it is used in protein synthesis. Lysine typically constitutes about 7-8% of an average protein. The epsilon-amino group often participates in hydrogen bonding and as a general base in catalysis. Common posttranslational modifications include methylation of the epsilon-amino group, giving methyl-, dimethyl-, and trimethyllysine. In bacteria, lysine is synthesized from aspartic acid, which is first converted to aspartyl-semialdehyde. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C6H14N2O2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 146.1876 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 146.105527702 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | KDXKERNSBIXSRK-YFKPBYRVSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C6H14N2O2/c7-4-2-1-3-5(8)6(9)10/h5H,1-4,7-8H2,(H,9,10)/t5-/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 56-87-1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2S)-2,6-diaminohexanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | L-lysine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NCCCC[C@H](N)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | L-alpha-amino acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 224.5 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Hydrogen ion + L-Lysine <> Cadaverine + Carbon dioxide Adenosine triphosphate + Water + L-Lysine > ADP + Hydrogen ion + L-Lysine + Phosphate Adenosine triphosphate + Water + L-Lysine > ADP + Hydrogen ion + L-Lysine + Phosphate Adenosine triphosphate + L-Lysine + tRNA(Lys) > Adenosine monophosphate + L-Lysine-tRNA (Lys) + Pyrophosphate Diaminopimelic acid + Hydrogen ion > Carbon dioxide + L-Lysine Fructoselysine-6-phosphate + Water <> Glucose 6-phosphate + L-Lysine Meso-2,6-Diaminoheptanedioate <> L-Lysine + Carbon dioxide L-Lysine <> Cadaverine + Carbon dioxide Adenosine triphosphate + L-Lysine + tRNA(Lys) + tRNA(Lys) <> Adenosine monophosphate + Pyrophosphate + L-Lysyl-tRNA + L-Lysyl-tRNA [tRNA(Ile2)]-cytidine34 + L-Lysine + Adenosine triphosphate <> [tRNA(Ile2)]-lysidine34 + Adenosine monophosphate + Pyrophosphate + Water -->-->Hydrogen ion + <i>meso</i>-diaminopimelate > L-Lysine + Carbon dioxide Adenosine triphosphate + L-Lysine + tRNA(Lys) > Adenosine monophosphate + Pyrophosphate + L-lysyl-tRNA(Lys) (tRNA(Ile2))-cytidine(34) + L-Lysine + Adenosine triphosphate > (tRNA(Ile2))-lysidine(34) + Adenosine monophosphate + Pyrophosphate + Water Biocytin + Water > Biotin + L-Lysine + L-Lysine Meso-2,6-Diaminoheptanedioate + Hydrogen ion > L-Lysine + Carbon dioxide + L-Lysine More...L-Lysine + Hydrogen ion + L-Lysine > Cadaverine + Carbon dioxide L-Lysine + L-Lysine > Cadaverine L-Lysine + Adenosine triphosphate + Hydrogen ion + tRNA(Lys) + L-Lysine > Adenosine monophosphate + Pyrophosphate + L-Lysyl-tRNA L-Lysine + Adenosine triphosphate + Water + L-Lysine > Adenosine diphosphate + Phosphate + Hydrogen ion + L-Lysine + ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Rothstein, Morton. DL-Lysine-6-C14 and DL-a-aminoadipic acid-6-C14. Biochemical Preparations (1961), 8 85-8. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in catalytic activity
- Specific function:
- Meso-2,6-diaminoheptanedioate = L-lysine + CO(2)
- Gene Name:
- lysA
- Locus Tag:
- PA5277
- Molecular weight:
- 45.5 kDa
Reactions
| Meso-2,6-diaminoheptanedioate = L-lysine + CO(2). |
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the binding-protein-dependent transport system for histidine. Probably responsible for energy coupling to the transport system
- Gene Name:
- hisP
- Locus Tag:
- PA2926
- Molecular weight:
- 28.5 kDa
- General function:
- Involved in nucleotide binding
- Specific function:
- ATP + L-lysine + tRNA(Lys) = AMP + diphosphate + L-lysyl-tRNA(Lys)
- Gene Name:
- lysS
- Locus Tag:
- PA3700
- Molecular weight:
- 57.3 kDa
Reactions
| ATP + L-lysine + tRNA(Lys) = AMP + diphosphate + L-lysyl-tRNA(Lys). |
- General function:
- Involved in nucleotide binding
- Specific function:
- With YjeK might be involved in the post-translational modification of elongation factor P (EF-P)
- Gene Name:
- poxA
- Locus Tag:
- PA5513
- Molecular weight:
- 32.4 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisM
- Locus Tag:
- PA2925
- Molecular weight:
- 26.7 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisQ
- Locus Tag:
- PA2924
- Molecular weight:
- 24.5 kDa
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the binding-protein-dependent transport system for histidine. Probably responsible for energy coupling to the transport system
- Gene Name:
- hisP
- Locus Tag:
- PA2926
- Molecular weight:
- 28.5 kDa
- General function:
- Involved in nucleotide binding
- Specific function:
- Probably part of a binding-protein-dependent transport system yecCS for an amino acid. Probably responsible for energy coupling to the transport system
- Gene Name:
- yecC
- Locus Tag:
- PA5152
- Molecular weight:
- 28.4 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisM
- Locus Tag:
- PA2925
- Molecular weight:
- 26.7 kDa
- General function:
- Involved in transport
- Specific function:
- Permease that is involved in the transport across the cytoplasmic membrane of lysine
- Gene Name:
- lysP
- Locus Tag:
- PA4628
- Molecular weight:
- 53.1 kDa
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for histidine; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- hisQ
- Locus Tag:
- PA2924
- Molecular weight:
- 24.5 kDa