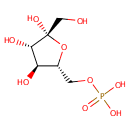
beta-D-Fructose 6-phosphate (PAMDB000547)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000547 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | beta-D-Fructose 6-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Beta-D-Fructose 6 phosphate (b-F6P) is the beta-anomer of fructose-6-phosphate. There are two anomers of fructose 6 phosphate, the alpha anomer and the beta anomer. Specifically, beta-D-fructose 6-phosphate is fructose sugar phosphorylated on carbon 6. Beta-D-Fructose 6-phosphate is a substrate for Fructose-1,6-bisphosphatase, Pyruvate kinase (isozymes R/L), Hexokinase (type I), Fructose-bisphosphate aldolase A, L-lactate dehydrogenase B chain, Glyceraldehyde-3-phosphate dehydrogenase and Transaldolase. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C6H13O9P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 260.1358 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 260.029718526 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | BGWGXPAPYGQALX-ARQDHWQXSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C6H13O9P/c7-2-6(10)5(9)4(8)3(15-6)1-14-16(11,12)13/h3-5,7-10H,1-2H2,(H2,11,12,13)/t3-,4-,5+,6-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | {[(2R,3S,4S,5R)-3,4,5-trihydroxy-5-(hydroxymethyl)oxolan-2-yl]methoxy}phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | β-D-fructofuranose 6-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | OC[C@@]1(O)O[C@H](COP(O)(O)=O)[C@@H](O)[C@@H]1O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as c-glycosyl compounds. These are glycoside in which a sugar group is bonded through one carbon to another group via a C-glycosidic bond. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carbohydrates and carbohydrate conjugates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Glycosyl compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | C-glycosyl compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + D-Fructose <> ADP + beta-D-Fructose 6-phosphate Mannose 6-phosphate <> beta-D-Fructose 6-phosphate Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate <> D-Erythrose 4-phosphate + beta-D-Fructose 6-phosphate beta-D-Fructose 6-phosphate + D-Glyceraldehyde 3-phosphate <> D-Erythrose 4-phosphate + Xylulose 5-phosphate Sorbitol-6-phosphate + NAD <> beta-D-Fructose 6-phosphate + NADH + Hydrogen ion beta-D-Glucose 6-phosphate <> beta-D-Fructose 6-phosphate Adenosine triphosphate + beta-D-Fructose <> ADP + beta-D-Fructose 6-phosphate Adenosine triphosphate + beta-D-Fructose 6-phosphate <> ADP + beta-D-Fructose 1,6-bisphosphate beta-D-Fructose 1,6-bisphosphate + Water <> beta-D-Fructose 6-phosphate + Phosphate D-Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate + D-Sedoheptulose 7-phosphate + D-Glyceraldehyde 3-phosphate <> beta-D-Fructose 6-phosphate + D-Erythrose 4-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||