Molybdopterin (PAMDB000446)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000446 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
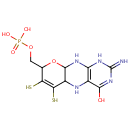
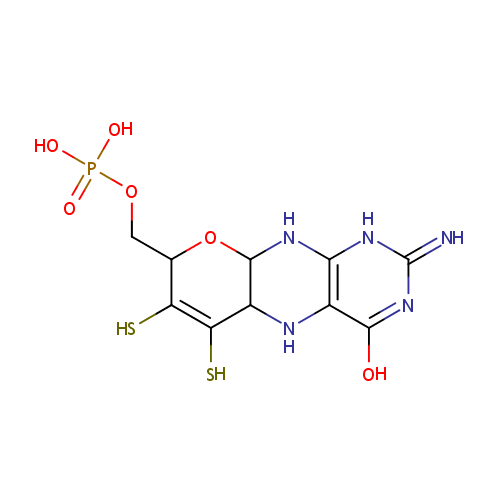
| Name: | Molybdopterin | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Molybdopterins are a class of biochemical cofactor that are used in many different enzymes. The simplest structure of molybdopterin contains a pyranopterin coordinated to molybdenum. The pyranopterin structure is a fused ring system containing a pyran fused to pterin. In addition, the pyran ring is substituted with two thiols and an alkyl phosphate. In molybdopterin, the thiols coordinate to molybdenum. In some cases, the alkyl phosphate group is replaced by an alkyl diphosphate nucleotide. -- Wikipedia | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H12MoN5O8PS2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 521.27 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 522.891898123 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | HDAJUGGARUFROU-UHFFFAOYSA-L | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H14N5O6PS2.Mo.2O/c11-10-14-7-4(8(16)15-10)12-3-6(24)5(23)2(21-9(3)13-7)1-20-22(17,18)19;;;/h2-3,9,12,23-24H,1H2,(H2,17,18,19)(H4,11,13,14,15,16);;;/q;+2;;/p-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 73508-07-3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | ({4-hydroxy-2-imino-6,7-disulfanyl-1H,2H,5H,5aH,8H,9aH,10H-pyrano[3,2-g]pteridin-8-yl}methoxy)phosphonic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | molybdopterin cofactor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NC1=NC2=C(NC3C(N2)OC(COP(O)(O)=O)C2=C3S[Mo](=O)(=O)S2)C(=O)N1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as molybdopterins. These are cofactors or analogs thereof, with a structure based on a furan ring fused to a pterin. In addition, the pyran ring features two thiolates, which serve as ligands in molybdo- and tungstoenzymes. In some cases, the alkyl phosphate group is replaced by an alkyl diphosphate nucleotide. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pteridines and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Pterins and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Molybdopterins | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Cyclic pyranopterin monophosphate + Copper + 2 MoaD Protein with thiocarboxylate >5 Hydrogen ion +2 MoaD Protein with carboxylate + Molybdopterin Guanosine triphosphate + Hydrogen ion + Molybdopterin > Molybdopterin guanine dinucleotide + Pyrophosphate Adenosine triphosphate + Hydrogen ion + Molybdopterin <> Adenylated molybdopterin + Pyrophosphate Molybdopterin + Adenylated molybdopterin > Adenosine monophosphate + bis-molybdenum cofactor + Copper 2 Hydrogen ion + Molybdate + Adenylated molybdopterin > Adenosine monophosphate + Copper + Water + Molybdopterin Cytidine triphosphate + Hydrogen ion + Molybdopterin > Molybdopterin cytosine dinucleotide + Pyrophosphate Cyclic pyranopterin monophosphate + 2 Sulfur donor <> Molybdopterin Adenosine triphosphate + Molybdopterin <> Pyrophosphate + Adenylated molybdopterin Cyclic pyranopterin monophosphate + Water + Thiocarboxylated-MPT-synthases > Molybdopterin + MPT-Synthases Cyclic pyranopterin monophosphate + Water + thiocarboxylated small subunit of molybdopterin synthase >4 Hydrogen ion +2 thiocarboxylated small subunit of molybdopterin synthase + Molybdopterin + Molybdopterin Molybdopterin + Adenosine triphosphate + Hydrogen ion + Molybdopterin > Pyrophosphate + Adenylyl-molybdopterin + Adenylyl-molybdopterin | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||