Aminoacetone (PAMDB000441)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000441 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
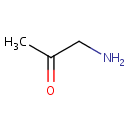
| Name: | Aminoacetone | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Threonine dehydrogenase catalyzes the oxidation of threonine by NAD+ to glycine and acetyl-CoA (5), but when the ratio acetyl-CoA/CoA increases in nutritional deprivation (e.g., in diabetes) the enzyme produces AA. (Chem. Res. Toxicol., 14 (9), 1323 -1329, 2001); | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
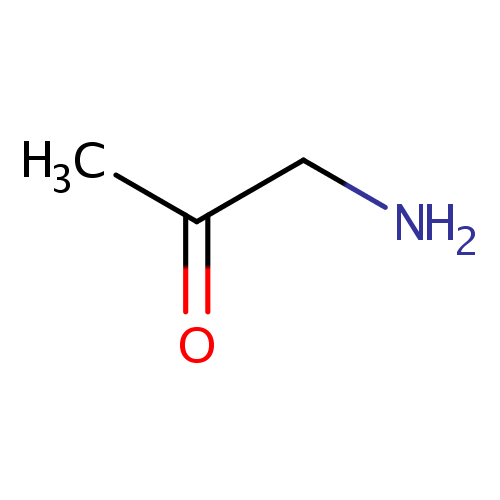
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C3H7NO | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 73.0938 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 73.052763851 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | BCDGQXUMWHRQCB-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C3H7NO/c1-3(5)2-4/h2,4H2,1H3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 298-08-8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 1-aminopropan-2-one | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | α-aminoacetone | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CC(=O)CN | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as alpha-amino ketones. These are ketones containing a carboxylic acid, and an amine group attached to the alpha carbon atom relative to C=O group. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carbonyl compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Ketones | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Alpha-amino ketones | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Aminoacetone + Hydrogen ion + NADH <> 1-Amino-2-propanol + NAD L-2-Amino-3-oxobutanoic acid + Hydrogen ion > Aminoacetone + Carbon dioxide Aminoacetone + Water + Oxygen <> Pyruvaldehyde + Ammonia + Hydrogen peroxide Aminoacetone + Water + Oxygen > Hydrogen ion + Pyruvaldehyde + Ammonia + Hydrogen peroxide L-Allothreonine + NADP + L-2-Amino-3-oxobutanoic acid <> Aminoacetone + Carbon dioxide + NADPH + Hydrogen ion 1-Amino-2-propanol + NAD + 4,5-Dihydro-4-hydroxy-5-S-glutathionyl-benzo[a]pyrene < NADH + Hydrogen ion + Aminoacetone | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Turner, J. M. Aminoacetone production by microorganisms. Biochemical Journal (1966), 98(1), 7P. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||