L-Homocysteine (PAMDB000067)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 1/22/2018 12:54:54 PM | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolite ID | PAMDB000067 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
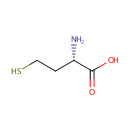
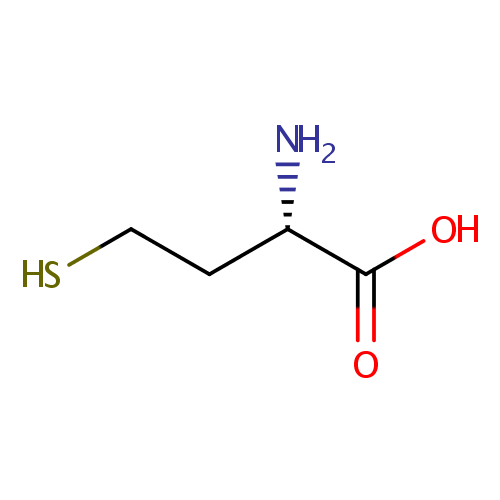
| Name: | L-Homocysteine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description: | Homocysteine is a thiol-containing amino acid formed by a demethylation of methionine. Homocysteine is a variant of the amino acid cysteine, differing in that its side-chain contains an additional methylene (-CH2-) group before the thiol (-SH) group. In Pseudomonas aeruginosa homocysteine (Hcy) is the last intermediate on the methionine biosynthetic pathway. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C4H9NO2S | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight: | 135.185 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight: | 135.035399227 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | FFFHZYDWPBMWHY-VKHMYHEASA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C4H9NO2S/c5-3(1-2-8)4(6)7/h3,8H,1-2,5H2,(H,6,7)/t3-/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 6027-13-0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2S)-2-amino-4-sulfanylbutanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | L-homocysteine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | N[C@@H](CCS)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Taxonomy Description | This compound belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | L-alpha-amino acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 232.6 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | L-Cystathionine + Water > L-Homocysteine + Ammonium + Pyruvic acid 5-Methyltetrahydrofolic acid + L-Homocysteine <> Hydrogen ion + L-Methionine + Tetrahydrofolic acid L-Homocysteine + S-Methylmethionine > Hydrogen ion +2 L-Methionine S-Adenosylmethionine + L-Homocysteine + S-Methylmethionine <> S-Adenosylhomocysteine + Hydrogen ion + L-Methionine S-Ribosyl-L-homocysteine > 4,5-Dihydroxy-2,3-pentanedione + L-Homocysteine S-Adenosylmethionine + L-Homocysteine <> S-Adenosylhomocysteine + L-Methionine 5-Methyltetrahydrofolic acid + L-Homocysteine <> Tetrahydrofolic acid + L-Methionine Cystathionine + Water <> L-Homocysteine + Ammonia + Pyruvic acid L-Cystathionine + Water <> L-Homocysteine + Ammonia + Pyruvic acid O-Succinyl-L-homoserine + Hydrogen sulfide <> L-Homocysteine + Succinic acid S-Ribosyl-L-homocysteine <> (4S)-4,5-Dihydroxypentan-2,3-dione + L-Homocysteine 5-Methyltetrahydropteroyltri-L-glutamic acid + L-Homocysteine <> Tetrahydropteroyltri-L-glutamic acid + L-Methionine L-Cystathionine + Water > Hydrogen ion + Pyruvic acid + Ammonia + L-Homocysteine L-Homocysteine + 5-Methyltetrahydropteroyltri-L-glutamic acid > L-Methionine + tetrahydropteroyl tri-L-glutamate L-Homocysteine + S-Adenosylmethionine Hydrogen ion + L-Methionine + S-Adenosylhomocysteine S-methyl-L-methionine + L-Homocysteine Hydrogen ion + L-Methionine 5-methyltetrahydropteroyltri-L-glutamate + L-Homocysteine > tetrahydropteroyltri-L-glutamate + L-Methionine L-Cystathionine + Water + 2-Aminoacrylic acid + 2-Iminopropanoate <> L-Homocysteine + Pyruvic acid + Ammonia | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Biochem Prepn. 5 93 (1957) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||